Montana Governor Gianforte just publicized a new DPHHS order requiring schools to provide a parental opt-out for mask requirements.
Underscoring the detrimental impact that universal masking may have on children, the rule cites a body of scientific literature that shows side effects and dangers from prolonged mask wearing.
The order purports to be evidence based. But is the evidence any good?
Mask Efficacy
The order cites:
The scientific literature is not conclusive on the extent of the impact of
masking on reducing the spread of viral infections. The department understands
that randomized control trials have not clearly demonstrated mask efficacy against
respiratory viruses, and observational studies are inconclusive on whether mask use
predicts lower infection rates, especially with respect to children.1
The supporting footnote is basically a dog’s breakfast,
1 See, e.g., Guerra, D. and Guerra, D., Mask mandate and use efficacy for COVID-19 containment in
US States, MedRX, Aug. 7, 2021, https://www.medrxiv.org/content/10.1101/2021.05.18.21257385v2
(“Randomized control trials have not clearly demonstrated mask efficacy against respiratory viruses,
and observational studies conflict on whether mask use predicts lower infection rates.”). Compare
CDC, Science Brief: Community Use of Cloth Masks to Control the Spread of SARS-CoV-2, last
updated May 7, 2021, https://www.cdc.gov/coronavirus/2019-ncov/science/science-briefs/masking-
science-sars-cov2.html, last visited Aug. 30, 2021 (mask wearing reduces new infections, citing
studies) ….
(more stuff of declining quality)
This is not an encouraging start; it’s blatant cherry picking. Guerra & Guerra is an observational statistical test of mask mandates. The statement DPHHS quotes, “Randomized control trials have not clearly demonstrated mask efficacy…” isn’t even part of the study; it’s merely an introductory remark in the abstract.
Much worse, G&G isn’t a “real” model. It’s just a cheap regression of growth rates against mask mandates, with almost no other controls. Specifically, it omits NPIs, weather, prior history of the epidemic in each state, and basically every other interesting covariate, except population density. It’s not even worth critiquing the bathtub statistics issues.
G&G finds no effects of mask mandates. But is that the whole story? No. Among the many covariates they omit is mask compliance. It turns out that matters, as you’d expect. From Leech et al. (one of many better studies DPHHS ignored):
Across these analyses, we find that an entire population wearing masks in public leads to a median reduction in the reproduction number R of 25.8%, with 95% of the medians between 22.2% and 30.9%. In our window of analysis, the median reduction in R associated with the wearing level observed in each region was 20.4% [2.0%, 23.3%]1. We do not find evidence that mandating mask-wearing reduces transmission. Our results suggest that mask-wearing is strongly affected by factors other than mandates.
We establish the effectiveness of mass mask-wearing, and highlight that wearing data, not mandate data, are necessary to infer this effect.
Meanwhile, the DPHHS downplays its second citation, the CDC Science Brief, which cites 65 separate papers, including a number of observational studies that are better than G&G. It concludes that masks work, by a variety of lines of evidence, including mechanistic studies, CFD simulations and laboratory experiments.
Verdict: Relying on a single underpowered, poorly designed regression to make sweeping conclusions about masks is poor practice. In effect, DPHHS has chosen the one earwax-flavored jellybean from a bag of more attractive choices.
Mask Safety
The department order goes on,
The department
understands, however, that there is a body of literature, scientific as well as
survey/anecdotal, on the negative health consequences that some individuals,
especially some children, experience as a result of prolonged mask wearing.2
The footnote refers to Kisielinski et al. – again, a single study in a sea of evidence. At least this time it’s a meta-analysis. But was it done right? I decided to spot check.
K et al. tabulate a variety of claims conveniently in Fig. 2:
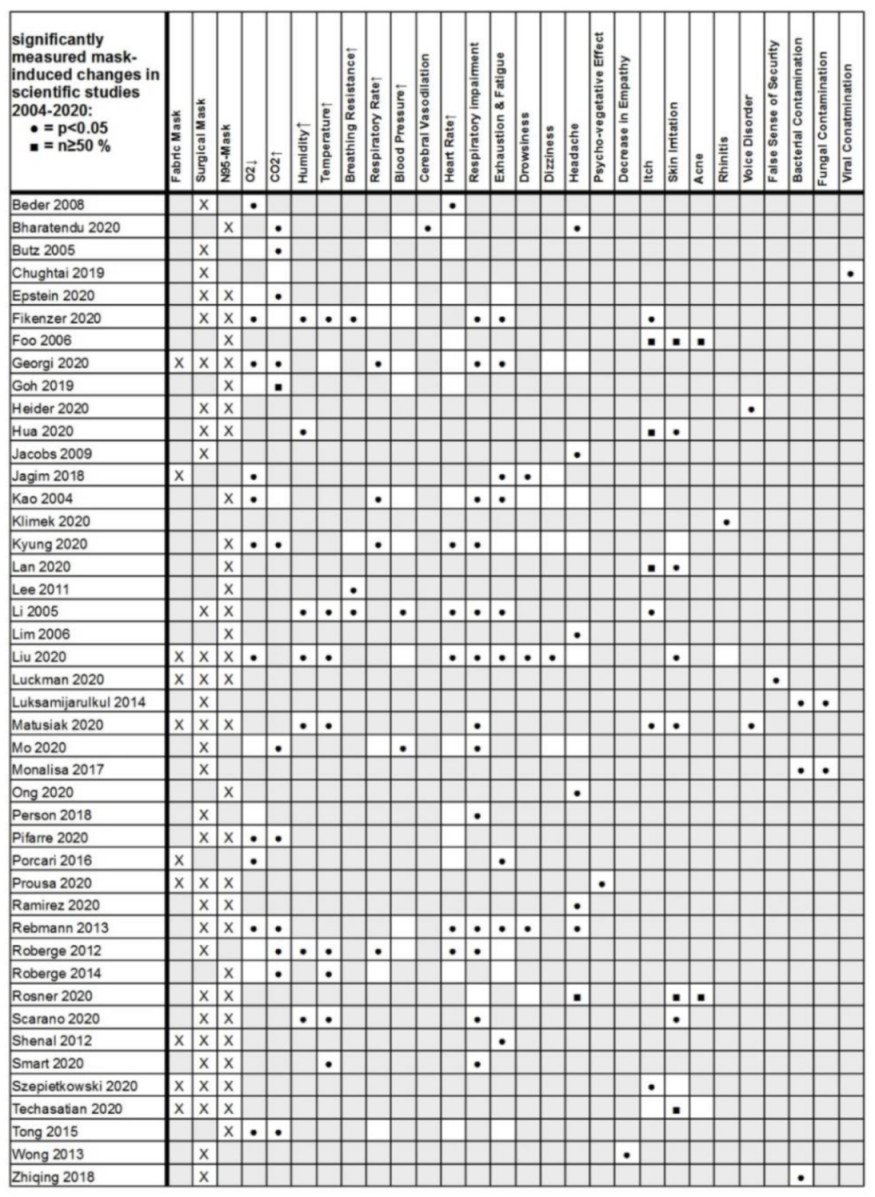
The first claim assessed is that masks reduce O2, so I followed those citations.
| Citation |
Claim |
Assessment/Notes |
| Beder 2008 |
Effect |
Effect, but you can’t draw any causal conclusion because there’s no control group. |
| Butz 2005 |
No effect |
PhD Thesis, not available for review |
| Epstein 2020 |
No effect |
No effect (during exercise) |
| Fikenzer 2020 |
Effect |
Effect |
| Georgi 2020 |
Effect |
Gray literature, not available for review |
| Goh 2019 |
No effect |
No effect; RCT n~=100 children |
| Jagim 2018 |
Effect |
Not relevant – this concerns a mask designed for elevation training, i.e. deliberately impeding O2 |
| Kao 2004 |
Effect |
Effect. End stage renal patients. |
| Kyung 2020 |
Effect |
Dead link. Flaky journal? COPD patients. |
| Liu 2020 |
Effect |
Small effect – <1% SpO2. Nonmedical conference paper, so dubious peer review. N=12. |
| Mo 2020 |
No effect |
No effect. Gray lit. COPD patients. |
| Person 2018 |
No effect |
No effect. 6 minute walking test. |
| Pifarre 2020 |
Effect |
Small effect. Tiny sample (n=8). Questionable control of order of test conditions. Exercise. |
| Porcari 2016 |
Effect |
Irrelevant – like Jagim, concerns an elevation training mask. |
| Rebmann 2013 |
Effect |
No effect. “There were no changes in nurses’ blood pressure, O2 levels, perceived comfort, perceived thermal comfort, or complaints of visual difficulties compared with baseline levels.” Also, no control, as in Beder. |
| Roberge 2012 |
No effect |
No effect. N=20. |
| Roberge 2014 |
No effect |
No effect. N=22. Pregnancy. |
| Tong 2015 |
Effect |
Effect. Exercise during regnancy. |
If there’s a pattern here, it’s lots of underpowered small sample studies with design defects. Morover, there are some blatant errors in assessment of relevance (Jagim, Porcari) and inclusion of uncontrolled studies (Beder, Rebmann, maybe Pifarre). In other words, this is 30% rubbish, and the rubbish is all on the “effect” side of the scale.
If the authors did a poor job assessing the studies they included, I also have to wonder whether they did a bad screening job. That turns out to be hard to determine without more time. But a quick search does reveal that there has been an explosion of interest in the topic, with a number of new studies in high-quality journals with better control designs. Regrettably, sample sizes still tend to be small, but the results are generally not kind to the assertions in the health order:
Mapelli et al. 2021:
Conclusions Protection masks are associated with significant but modest worsening of spirometry and cardiorespiratory parameters at rest and peak exercise. The effect is driven by a ventilation reduction due to an increased airflow resistance. However, since exercise ventilatory limitation is far from being reached, their use is safe even during maximal exercise, with a slight reduction in performance.
Chan, Li & Hirsch 2020:
In this small crossover study, wearing a 3-layer nonmedical face mask was not associated with a decline in oxygen saturation in older participants. Limitations included the exclusion of patients who were unable to wear a mask for medical reasons, investigation of 1 type of mask only, Spo2 measurements during minimal physical activity, and a small sample size. These results do not support claims that wearing nonmedical face masks in community settings is unsafe.
Lubrano et al. 2021:
This cohort study among infants and young children in Italy found that the use of facial masks was not associated with significant changes in Sao2 or Petco2, including among children aged 24 months and younger.
Shein et al. 2021:
The risk of pathologic gas exchange impairment with cloth masks and surgical masks is near-zero in the general adult population.
A quick trip to PubMed or Google Scholar provides many more.
Verdict: a sloppy meta-analysis is garbage-in, garbage-out.
Bottom Line
Montana DPHHS has failed to verify its sources, ignores recent literature and therefore relies on far less than the best available science in the construction of its flawed order. Its sloppy work will fan the flames of culture-war conspiracies and endanger the health of Montanans.